DNA-seq_Somatic_Variants
Somatic Variants Pipeline
This repository is designed to analyze somatic mutations from HCC1143 tumor samples with a matched normal control. The pipeline includes preprocessing, somatic variant calling, filtering, annotation, and copy number variation (CNV) detection using widely used bioinformatics tools.
Table of Contents
- Server Information
- Genomic Sequencing Data
- Preprocessing Steps
- Somatic Variant Calling and Filtering
- Annotate Variants
- Copy Number Variation
1. Server Information
Prerequisites
- Tools: Install PuTTY, FileZilla, IGV
- Server Details:
- IP Address:
168.105.161.70 - Port:
22 - Access: Requires JABSOM or UH network (use VPN for remote access).
- Note: With PuTTY and FileZilla you can connect to server.
- IP Address:
Security Practices
- Avoid multiple failed login attempts to prevent account locking.
- Use strong passwords or SSH keys.
- Log out after completing tasks.
Installing Miniconda
wget https://repo.continuum.io/miniconda/Miniconda3-latest-Linux-x86_64.sh
bash Miniconda3-latest-Linux-x86_64.sh
source ~/.bashrc
conda config --add channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge
Installing software
# conda install mamba
# mamba install sra-tools fastqc trimmomatic multiqc curl
# mamba install bwa samtools picard gatk4
# mamba install snpeff snpsift
2. Genomic Sequencing Data
Sample Information
- Sample: HCC1143 (Breast Cancer Cell Line)
- Matched Normal Sample: HCC1143_normal
- Data files:
- Tumor BAM:
/home/bqhs/mutect2/tumor.bam - Normal BAM:
/home/bqhs/mutect2/normal.bam - Reference Genome:
/home/bqhs/mutect2/Homo_sapiens_assembly38.fasta - Panel of Normals (PoN):
/home/bqhs/mutect2/chr17_m2pon.vcf.gz - Germline Resource:
/home/bqhs/mutect2/chr17_af-only-gnomad_grch38.vcf.gz - Target Intervals:
/home/bqhs/mutect2/targets_chr17.interval_list
- Tumor BAM:
- In the interest of time,somatic variant calling analysis is restricted to chromosome 17
- Common biallelic SNPs: /home/bqhs/mutect2/chr17_small_exac_common_3_grch38.vcf.gz (for GetPileupSummaries)
3. Preprocessing Steps
Steps Common to Germline and Somatic Variants
- Quality Control:
- FastQC, Trimmomatic → Checks read quality and trims adapters.
- Read Alignment:
- bwa mem → Maps reads to the reference genome.
- Sort alignment
- samtools sort → Organizes BAM file
- Mark duplicates
- picard MarkDuplicates → Removes PCR artifacts
- Recalibrate base quality scores
- GATK BaseRecalibrator & GATK ApplyBQSR → Adjusts systematic sequencing errors
- Index BAM file
- samtools index → Enables efficient access to BAM files.
4. Somatic Variant Calling and Filtering
mkdir -p Somatic_mutation/Exercise
cd Somatic_mutation/Exercise
1. Run GATK Mutect2
- GATK Mutect2 is a variant caller used to detect somatic mutations in cancer samples.
- It compares sequencing data from a tumor sample to a matched normal sample (if available) to identify mutations unique to the tumor.
- Arguments:
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta- Reference genome (GRCh38) used to align reads and determine variant locations.
-I /home/bqhs/mutect2/tumor.bam- Input tumor BAM file (aligned reads from the cancer sample).
-tumor HCC1143_tumor- Specifies the tumor sample name in the BAM file.
-I /home/bqhs/mutect2/normal.bam- Input normal BAM file (optional) – helps filter out germline mutations.
-normal HCC1143_normal- Specifies the normal sample name in the BAM file.
-pon /home/bqhs/mutect2/chr17_m2pon.vcf.gz- Panel of Normals (PoN), a database of common sequencing artifacts (not real mutations) to avoid false positives.
--germline-resource /home/bqhs/mutect2/chr17_af-only-gnomad_grch38.vcf.gz- Germline variant database (gnomAD) used to filter out inherited variants so only tumor-specific mutations remain.
-L /home/bqhs/mutect2/chr17plus.interval_list- Restricts analysis to specific regions (in this case, chromosome 17) to save time and focus on key areas.
-O somatic_m2.vcf.gz- Output VCF file where the detected somatic variants are stored.
gatk Mutect2 \
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
-I /home/bqhs/mutect2/tumor.bam -tumor HCC1143_tumor \
-I /home/bqhs/mutect2/normal.bam -normal HCC1143_normal \
-pon /home/bqhs/mutect2/chr17_m2pon.vcf.gz \
--germline-resource /home/bqhs/mutect2/chr17_af-only-gnomad_grch38.vcf.gz \
-L /home/bqhs/mutect2/chr17plus.interval_list \
-O somatic_m2.vcf.gz
Extract Read Group Information from BAM Header
samtools view -H /home/bqhs/mutect2/tumor.bam | grep '@RG'
samtools view -H /home/bqhs/mutect2/normal.bam | grep '@RG'
Extract Sample Name Using GATK
gatk GetSampleName -I /home/bqhs/mutect2/tumor.bam -O tumor.txt
gatk GetSampleName -I /home/bqhs/mutect2/normal.bam -O normal.txt
Extract subsets records that contain a comma in the 5th column.
zcat somatic_m2.vcf.gz | awk '$5 ~","'
https://gatk.broadinstitute.org/hc/en-us/articles/360035531692-VCF-Variant-Call-Format
https://gatk.broadinstitute.org/hc/en-us/articles/360035531912-Spanning-or-overlapping-deletions-allele-
Displays the first 10,000 lines of the decompressed VCF file
zcat somatic_m2.vcf.gz | head -n 10000
View the standard Format in the VCF header
zcat somatic_m2.vcf.gz | grep '##FORMAT'
View the standard INFO in the VCF header
zcat somatic_m2.vcf.gz | grep '##INFO'
2. Run GATK GetPileupSummaries
- Runs GATK’s GetPileupSummaries tool to compute pileup summaries for given sites.
- This command is used to summarize allele frequencies at common germline variant sites
- Arguments:
-R Homo_sapiens_assembly38.fasta- Reference genome file (GRCh38) required for alignment consistency.
-I Input BAM file (normal sample)- containing aligned sequencing reads.
-V chr17_small_exac_common_3_grch38.vcf.gz- A population germline variant VCF (from gnomAD/ExAC), which contains known common variants. This is used to distinguish somatic from germline variants.
-L targets_chr17.interval_list- Restricts analysis to specific genomic regions (e.g., targeted exome or panel).
O normal.pileups.table- Output file storing pileup summaries, including:
- Chromosome, position, reference allele
- Counts for reference and alternative alleles
- Population allele frequency estimates
- Output file storing pileup summaries, including:
- This step is preparation for contamination estimation, which is crucial for Mutect2 variant calling.
- Helps detect tumor-normal contamination by analyzing allele frequencies in the normal sample.
# For Normal
gatk GetPileupSummaries \
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
-I /home/bqhs/mutect2/normal.bam \
-V /home/bqhs/mutect2/chr17_small_exac_common_3_grch38.vcf.gz \
-L /home/bqhs/mutect2/targets_chr17.interval_list \
-O normal.pileups.table
# For Tumor
gatk GetPileupSummaries \
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
-I /home/bqhs/mutect2/tumor.bam \
-V /home/bqhs/mutect2/chr17_small_exac_common_3_grch38.vcf.gz \
-L /home/bqhs/mutect2/targets_chr17.interval_list \
-O tumor.pileups.table
Displays the contents of the tumor pileup summary table.
- Filters for lines that start with “chr17”
- Uses awk to check if the fifth column (alternate allele count) is greater than or equal to 3.
- The 5th column represents the number of alternate allele supporting reads at a given site. This filters sites where at least 3 reads support an alternate allele.
cat tumor.pileups.table | grep '^chr17' | awk '$5>=3'
cat normal.pileups.table | grep '^chr17' | awk '$5>=3'
3. Run GATK CalculateContamination
- This estimates tumor sample contamination using allele frequencies from the pileup summaries.
- Arguments:
-I tumor.pileups.table- Input pileup summary for the tumor sample, which contains allele counts at known germline variant sites.
-matched normal.pileups.table- The corresponding normal sample pileup summary, used to refine contamination estimation.
-O contamination.table- Output file storing contamination estimates.
- If no matched normal is available, contamination can still be estimated, but with reduced accuracy.
gatk CalculateContamination \
-I tumor.pileups.table \
-matched normal.pileups.table \
-O contamination.table
4. Run GATK FilterMutectCalls
- Filter somatic variants → Removes low-confidence calls
- Removes false positives caused by sequencing errors, contamination, and other artifacts.
- Arguments:
-V somatic_m2.vcf.gz- Input VCF file generated by Mutect2, containing raw somatic variants (before filtering).
--contamination-table contamination.table- Provides contamination estimates from CalculateContamination to filter out potential contaminant alleles.
-R Homo_sapiens_assembly38.fasta- Reference genome (GRCh38) used for consistency.
- `-O somatic.filtered.vcf.gz
- Output filtered VCF file, containing high-confidence somatic mutations
gatk FilterMutectCalls \
-V somatic_m2.vcf.gz \
--contamination-table contamination.table \
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
-O somatic.filtered.vcf.gz
# contamination-table: Tables containing contamination information
zcat somatic.filtered.vcf.gz | grep '##FILTER'
zcat somatic.filtered.vcf.gz | grep 'germ-line'
zcat somatic.filtered.vcf.gz | grep 'PASS'
zcat somatic.filtered.vcf.gz | grep 'contamiation'
5. Run GATK CollectSequencingArtifactMetrics
- A tool from GATK’s Picard suite that analyzes systematic sequencing errors in BAM files.
- Arguments:
-R Homo_sapiens_assembly38.fasta- The reference genome (GRCh38) used for alignment.
-I tumor.bam- Input BAM file containing aligned reads for the tumor sample.
EXT ".txt"- Specifies the file extension for output reports (default is .metrics, but here it’s set to .txt).
-O tumor_artifact- Prefix for the output artifact metrics files.
gatk CollectSequencingArtifactMetrics \
-R /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
-I /home/bqhs/mutect2/tumor.bam \
-EXT ".txt" \
-O tumor_artifact
5. Annotate Variants
Functional annotation
- This steps functionally annotates somatic variants using SnpEff and SnpSift.
- SnpEff for variant annotation
- SnpSift for variant filtering and post-processing of annotated VCF files
SnpEff does not accept compressed VCF files. So unzip the file.
gunzip -c somatic.filtered.vcf.gz > somatic.filtered.vcf
snpEff is a variant effect predictor that annotates variants with their potential functional impact.
- Input: somatic.filtered.vcf (filtered somatic variants).
- Output: somatic.filtered.ann.vcf (annotated VCF file).
- Arguments:
-Xmx2gAllocates 2GB of memory for Java (adjust if needed).annAnnotation mode.hg38Reference genome database (ensure it is installed in SnpEff).-vVerbose mode (detailed logs).-s snpeff.htmlGenerates a summary report (snpeff.html), which includes: Variant effects, Gene and protein impact predictions
snpEff -Xmx2g ann hg38 -v -s snpeff.html somatic.filtered.vcf > somatic.filtered.ann.vcf
Annotate with dbSNP Using SnpSift
- Input annotated VCF.
- Output VCF, now with dbSNP annotations.
- Adds known dbSNP IDs (rsIDs) to the VCF.
/home/bqhs/hg38/dbsnp_146.hg38.vcf.gzReference dbSNP database (version 146, built for hg38).
SnpSift annotate /home/bqhs/hg38/dbsnp_146.hg38.vcf.gz somatic.filtered.ann.vcf > somatic.filtered_1.ann.vcf
Lists sample names present in the final annotated VCF
# mamba install bcftools
bcftools query -l somatic.filtered_1.ann.vcf
- SnpEff: Predicts functional impact (e.g., missense, nonsense, synonymous).
- SnpSift: Adds dbSNP rsIDs for variant reference.
- bcftools query: Ensures sample metadata is intact.
Extract relevant fields
- SnpSift extractFields → Filters and formats VCF for downstream analysis
SnpSift extractFields somatic.filtered_1.ann.vcf \ ID CHROM POS REF ALT QUAL DP FILTER \ ANN[0].GENE ANN[0].GENEID ANN[0].EFFECT ANN[0].IMPACT \ ANN[0].BIOTYPE ANN[0].HGVS_C ANN[0].HGVS_P \ GEN[0].GT GEN[0].GQ GEN[0].FT \ GEN[1].GT GEN[1].GQ GEN[1].FT > somatic_final.txt
6. Copy Number Variation
Create a New Conda Environment with Python 3.9
conda create -n cnvkit_env python=3.9 -y
Activate the New Environment
conda activate cnvkit_env
Install Pandas 1.5.3
conda install pandas=1.5.3 -y
Install CNVkit
conda install -c bioconda cnvkit
Verify CNVkit Installation
cnvkit.py version
Run CNV detection using CNVKit
cnvkit.py batch→ Detects large deletions/amplifications- This CNVkit command processes somatic copy number variations (CNVs) in a tumor-normal paired analysis. It generates CNV profiles for the tumor sample using a matched normal sample for comparison.
- Starting files:
- Use the same BAM files and reference genome as Mutect2 exercise
- Target list: /home/bqhs/mutect2/targets_chr17.interval_list
- Gene annotation: /home/bqhs/mutect2/refFlat.txt
- Observe the effect of changing the purity setting when converting to copy number (integers)
cnvkit.py batch /home/bqhs/mutect2/tumor.bam \
-n /home/bqhs/mutect2/normal.bam \
-t /home/bqhs/mutect2/targets_chr17.interval_list \
-f /home/bqhs/mutect2/Homo_sapiens_assembly38.fasta \
--annotate /home/bqhs/mutect2/refFlat.txt \
--scatter --diagram \
-d cnvkit_output
Call CNVs
- (cnvkit.py call) → Determines CNV events per sample
- This command calls copy number alterations (CNAs) from the segmented CNV data (tumor.cns). It converts log2 ratio values into discrete copy number states (e.g., deletions, amplifications, and neutral regions).
- CNVkit infers integer copy numbers from the log2 ratio data in tumor.cns.
- It classifies segments as:
- Loss (CN < 2, e.g., deletions)
- Gain (CN > 2, e.g., amplifications)
- Neutral (CN = 2, normal diploid regions)
cd cnvkit_output
cnvkit.py call tumor.cns -o tumor.call.cns
Copy Number Variation (CNV) plot for a tumor sample
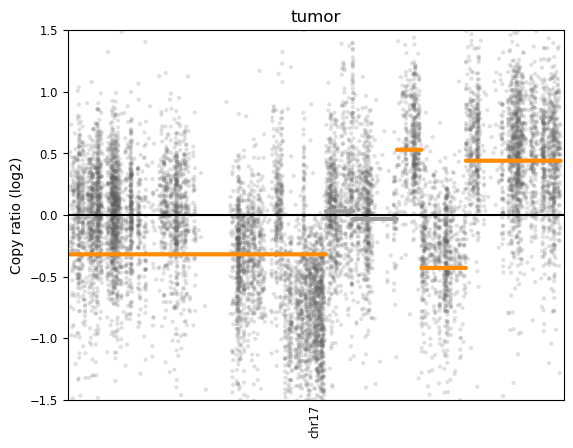
- X-axis (chr17): Represents the genomic position along chromosome 17.
- Y-axis (Copy Ratio log2): Represents the log2-transformed copy number ratio, where:
- 0 (baseline) indicates a normal diploid copy number.
- Values above 0 indicate copy number gains (amplifications).
- Values below 0 indicate copy number losses (deletions).
- Gray Dots: Represent individual data points for copy number variations across the chromosome.
- Orange Segments: Likely represent significant CNV regions, identified by segmentation algorithms.
Deactivate Environment After Running
conda deactivate
Few more steps
Examine results in IGV.
- Copy few files and save them in your local machine.
cp /home/bqhs/mutect2/tumor.bam ./
cp /home/bqhs/mutect2/normal.bam ./
samtools index tumor.bam
samtools index normal.bam
cp /home/bqhs/mutect2/chr17_m2pon.vcf.gz ./
cp /home/bqhs/mutect2/chr17_af-only-gnomad_grch38.vcf.gz ./
- Load:
- The BAM files
- Germline AF resource and PoN VCF
- Your filtered Mutect2 results (VCF)
- What do you observe at these sites?
- chr17:7,666,402-7,689,550